In meiosis, maternal and paternal chromosomes mutually exchange some parts of their chromosome arms to form genetically diverse generative cells. The process that shuffles genetic traits is called meiotic homologous recombination. However, recombination of repetitive DNA, for instance the ribosomal DNA (rDNA) clusters, can be dangerous, ultimately leading to loss of rDNA repeats. Ribosomal DNA encodes the information of the RNA share of ribosomes. Ribosomes, gigantic complexes comprised of RNA and proteins are the protein factories of a cell. Loss of rDNA units is detrimental to the fitness of an organism because it decreases the capacity to produce new proteins. These are needed all the time, but especially during developmental transitions or stress conditions.
Unlike the more prominent homologous recombination (HR) mechanism, the non-homologous end joining mechanism (NHEJ) repairs DSBs by simply “stitching” the ends of DNA together. NHEJ has been thought to play no substantial role during meiosis, a view challenged by the findings of the Schlögelhofer lab. It turns out that NHEJ has an important function in repairing rDNA. Utilizing the model plant Arabidopsis thaliana, the researchers found that early in meiosis all of the hundreds of rDNA repeats localize in the nucleolus. There they are shielded from acquiring a meiosis-specific chromatin signature and from the canonical meiotic DSB machinery. Lesions within the rDNA are repaired via NHEJ, and not via HR, revealing an important role of NHEJ in maintaining the integrity of the rDNA over many generations.
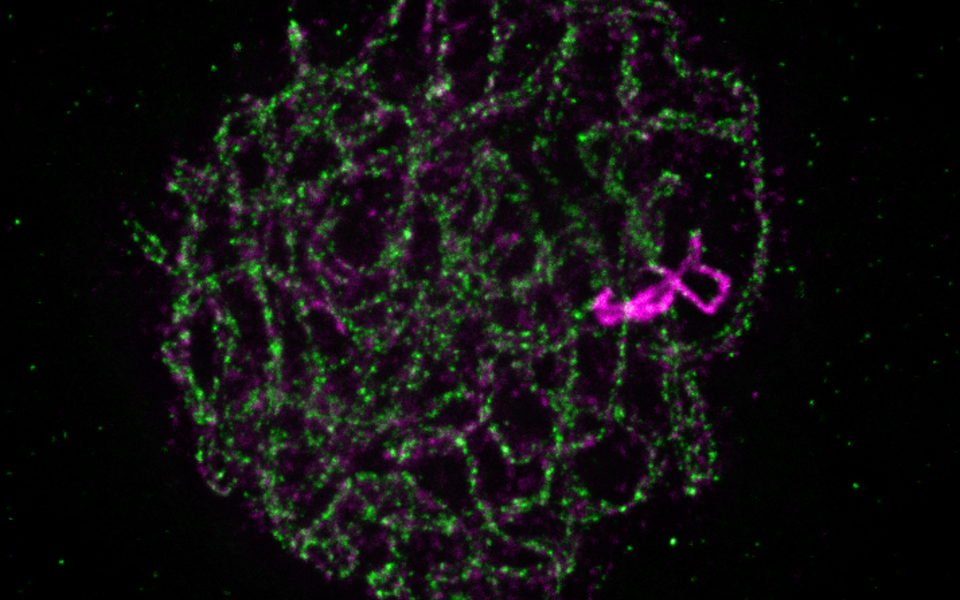
Developing novel methods and the latest microscopy technologies played an important role in the research. The scientists managed to visualize DNA and proteins in a 3D environment, as first author Jason Sims describes: “Imaging cells, nuclei and DNA in their natural spatial contexts and configurations is challenging yet required to understand the biology of genome organization, maintenance and transmission. Here we developed an advanced “whole mount immuno-FISH” (WhoMI-FISH) method, specifically optimized for pollen mother cells (PMCs) of Arabidopsis thaliana. It allows specimen preparation that maintains meiocyte nuclei positions and genome organization and simultaneous detection of specific genomic regions and meiotic proteins.”
Original Publication in The Plant Cell:
Jason Sims, Gregory P. Copenhaver, Peter Schlögelhofer: Meiotic DNA Repair in the Nucleolus Employs a Non-homologous End Joining Mechanism.